Catalog Number|Packaging
Mat. No |
Ref. No |
No. of preps |
|
4992901 |
GKM101 | 20 rxn |
-
Features
● Simple and fast: Only four steps are required to obtain mutants. Unlike traditional strand displacement amplification (SDA), this kit does not require multi-round PCR or the construction of sub-clones.
● High-efficiency primers: Part-overlapping primers were designed to generate more desired mutated plasmids by PCR amplification.
● Wide applications: Both single and multiple mutations can be applied by using this kit, and the mutation sites can be up to five within a single plasmid.
● High suitability: Size of the target plasmid can be up to 10 kb, which means almost all the common used plasmids suit to this kit.
● High mutation rate: Methylated plasmid can be digested both in vivo and in vitro by using this kit, ensuring higher mutation rate. And for single mutation, the mutation rate is higher than 90%.
-
Description
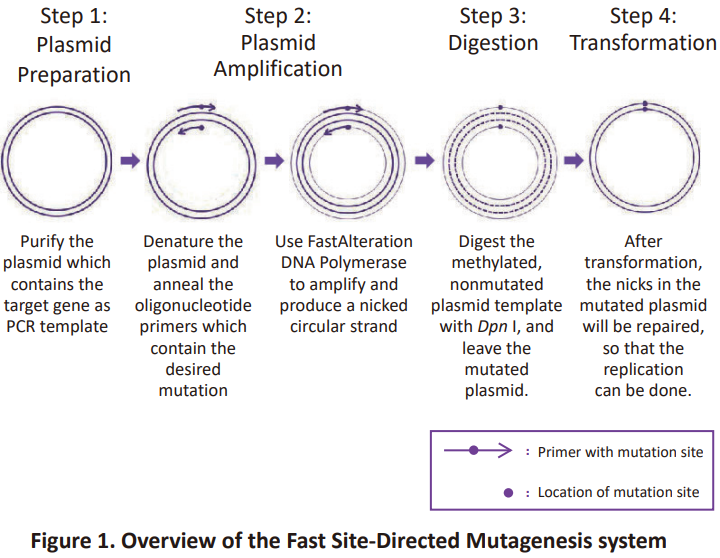
In vitro site-directed mutagenesis is a crucial technique for the modification and optimization of target gene, studying regulatory site of promoter and the complicated relationship between protein’s structure and function. The Fast Site-Directed Mutagenesis Kit allows site-specific mutation at target gene, including single and multiple mutations, insertion or deletion mutations. The Fast Site-Directed Mutagenesis Kit can promise a 90% or even higher mutation rate for introducing a single mutation to target gene. Different from previous low efficient mutagenesis kit, this Fast Site-Directed Mutagenesis Kit does not require multi-round PCR or sub-clones which cost lots of time and labor; only four steps are required to construct mutants (Figure 1).
FastAlteration DNA Polymerase in this kit is a DNA polymerase with high fidelity, high speed and high sensitivity. It could amplify up to 10 kb of plasmid at 15-30 sec/kb. FDM competent cells could digest methylated plasmid within the cell, it means that they could degrade those plasmids which are failed to be degraded by Dpn I and then ensure a higher positive rate. Meantime, this kit also contains control plasmid and primers to help customers to locate experimental problems.
-
Kit Contents
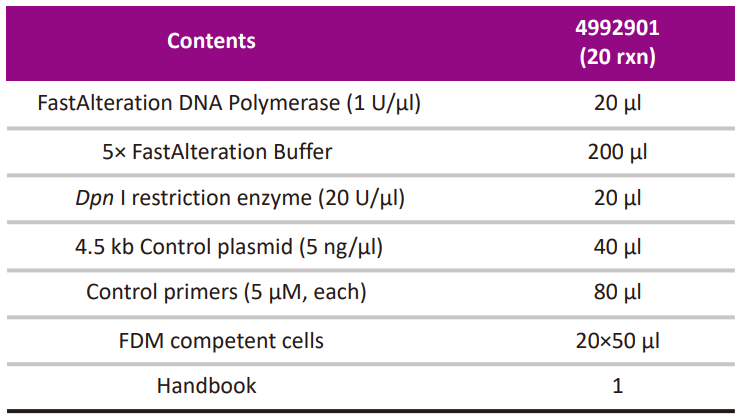
-
Applications
This kit could be used to optimize the target gene or vector, analyze the sites on promoter which interact with regulatory proteins, and study the relationship between the structure and function of proteins. -
Storage Condition
● FDM competent cells: Please store the FDM competent cells at -90~-65°C right after you receive it. Cells would be stable for 6 months at -90~-65°C.
● All other components should be stored at -30~-15°C, at which they are stable for 15 months.
-
Important Notes
● When multiple mutations are applied on a single primer, the more mutation sites it contains, the lower mutation rate occurs. Previous data suggested that when five mutation sites were designed on a single primer, the mutation rate were around 50%. We suggest increasing the clones number to verify.
● This kit allows multiple mutations in multiple primers, which gives a wider range for mutation study in a certain gene. The upper limit of mutation sites is still five.
● Control plasmid and primers are strongly recommended to be used for the troubleshooting reason.
-
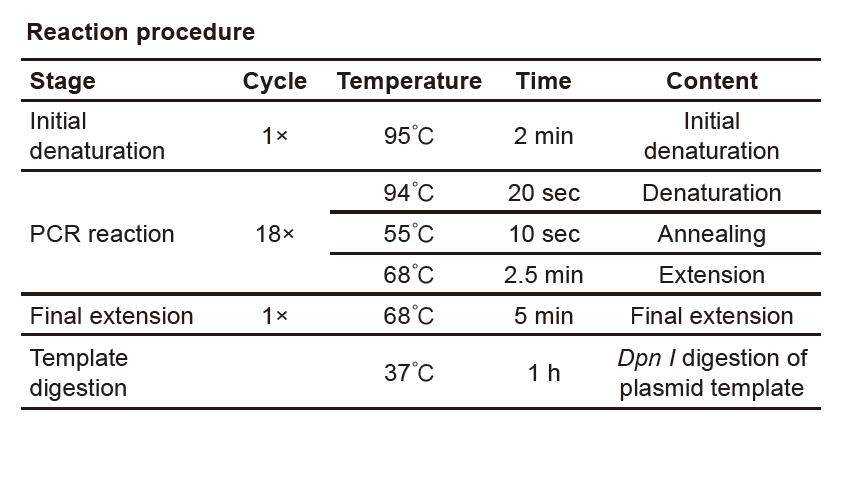
Site-Mutation Reaction Setup and PCR Program
Note: The reaction mentions below is just an example case, the real reaction condition should depend on the specific circumstance.
● Thoroughly thaw and shake the template plasmid, mutagenic primer solutions and all the other PCR reagents.
● Prepare a reaction solution according to the following table:
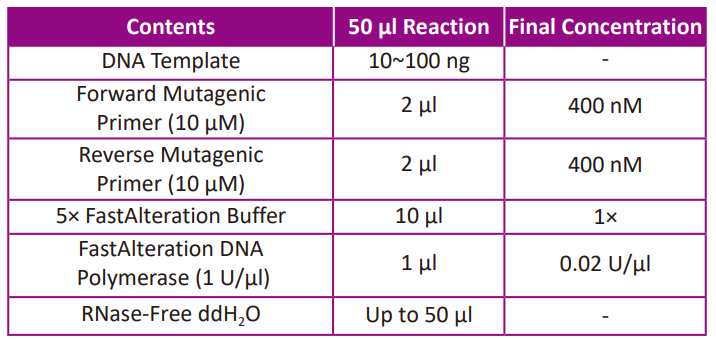
● Prepare the control PCR reaction according to the following table.
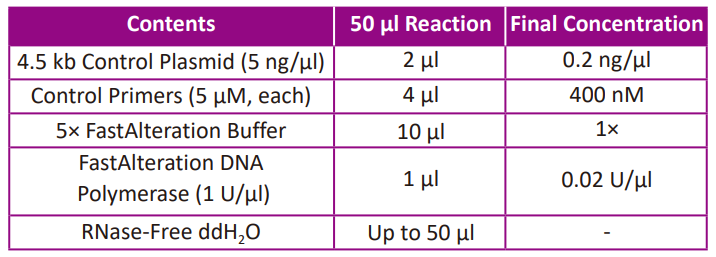
● Start the PCR program showed below for the two reactions.
Note: The PCR program settings could be changed depending on the specific circumstances.
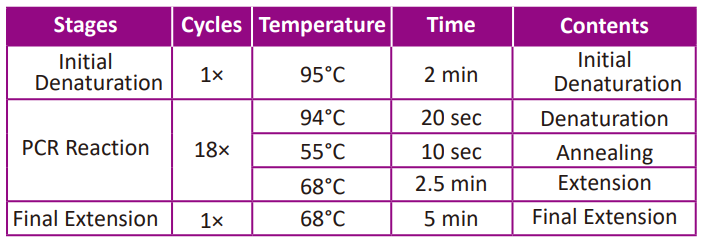
-
Principles of Primer Design
● Both of the mutagenic primers must have the desired mutation sites on them, and the primers’ sequences should be complementary with the target plasmid except for the mutation site.
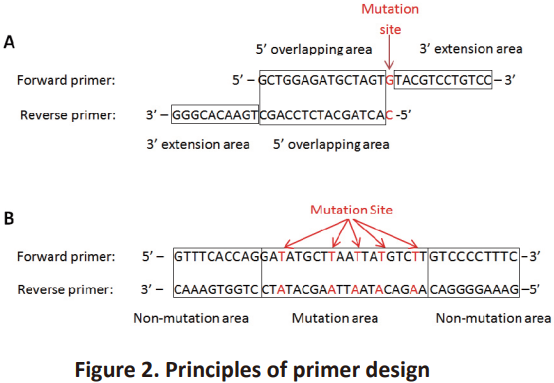
● If there is only one mutation site in a primer, it should be designed as what is showed in Figure 2-A. This kind of primer should contain two parts: 5’ overlapping area and 3’ extension area. Primers should be around 30 bases in length which include 15-20 bases of 5’ overlapping area and at least 10 bases of 3’ extension area. The mutation site should locate at the downstream of forward primer’s overlapping area and at the 5’ end of reverse primer.
● If there are 2-5 mutation sites in a primer, it should be designed as what is showed in Figure 2-B. The sequences of this primer pair are completely complementary and contain two parts: mutation area and non-mutation area. The length of primer should be around 40 bases, within which the mutation area should be 15-20 bases and the non-mutation area should be at least 10 bases. 2-5 mutation sites could be applied depending on the experimental purpose.
● Primers must be purified either by high performance liquid chromatography (HPLC) or by polyacrylamide gel electrophoresis (PAGE).
● The following formula is commonly used for estimating the Tm of primers:
Tm = 81.5 + 0.41(%GC) − 675/N − mismatch%
(N stands for the length of primer)
-
Sort by
-
Date
Date(
)
Date
Date(
)
Impact Factor
IF(
)
Impact Factor
IF(
)



 Inquire
Inquire