Catalog Number|Packaging
Mat. No |
Ref. No |
No. of preps |
-
Description
Bacterial and fungal whole genome re-sequencing is a critical tool to complete the genomes of known bacterium and fungi, includes third generation sequencing, assembly, functional annotation and advanced bioinformatic analysis fulfilling specific research goals. A more comprehensive profiling of bacteria genome empowers revealing of fundamental mechanisms underlying their biological processes, which could also provide valuable reference for genomic researches in higher eukaryotic species, etc.
-
Technical Workflow
Scheme design → Sample processing → Library construction sequencing → Data analysis → After-sales service
-
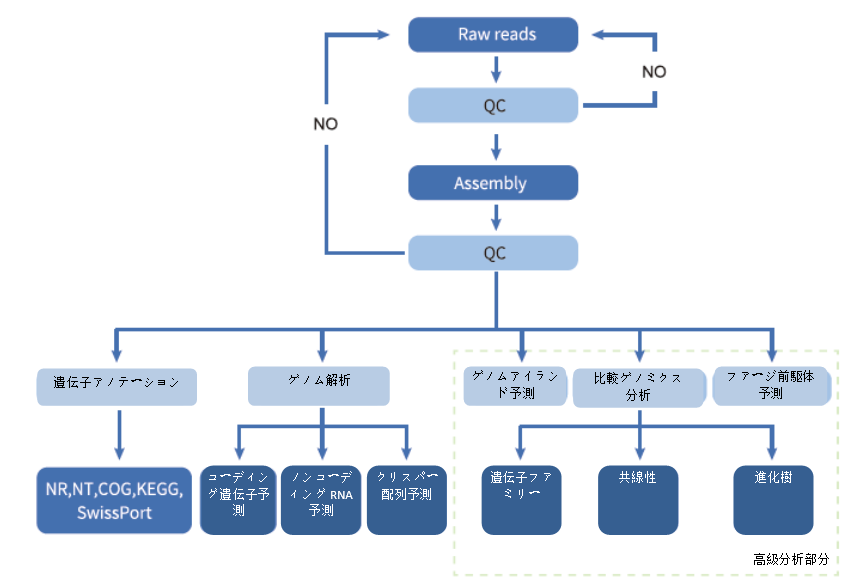
Analysis Workflow
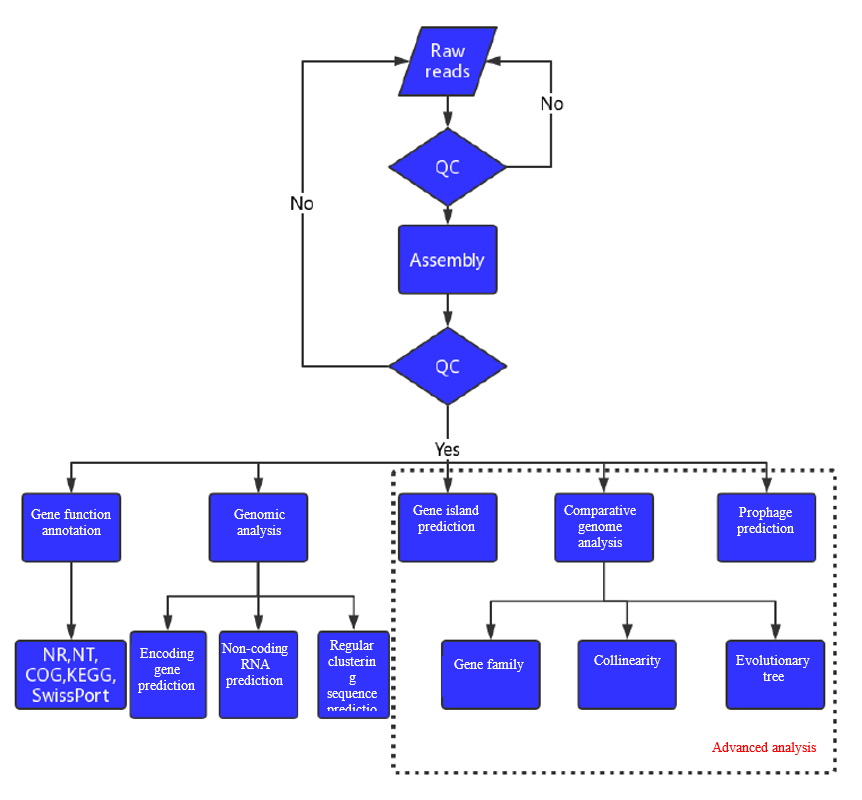
-
Features
1. As a global supplier of high-quality NGS raw materials, we guarantee the quality of the project from the source, including high-quality extraction and library construction ect.;
2. Senior technical teams, with more than ten years of information analysis experience, can provide a variety of personalized analysis;
3. With high-quality after-sales service, no longer need to worry about the results anymore;
-
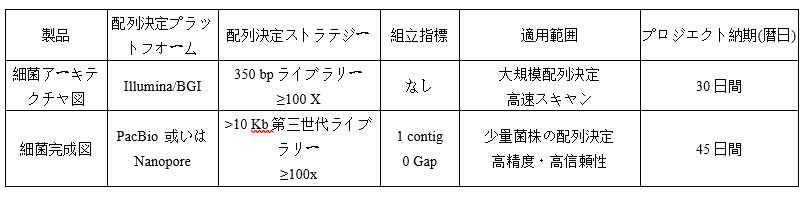
Parameter
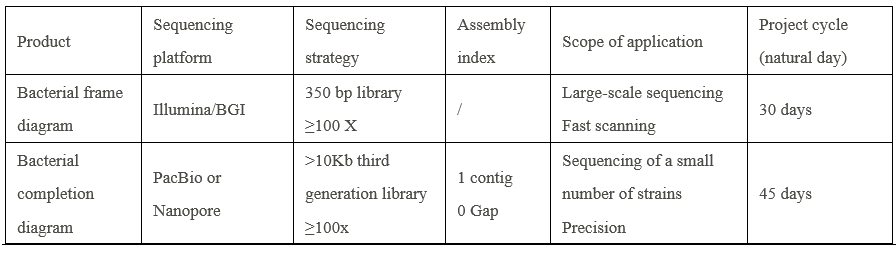
-
Applications
Industrial field, human microbiology, environmental microbiology, agricultural field
-
Requirements of Samples
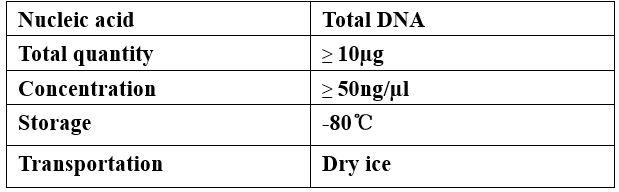
-
Results Demo
Circos diagram of gene and annotation results
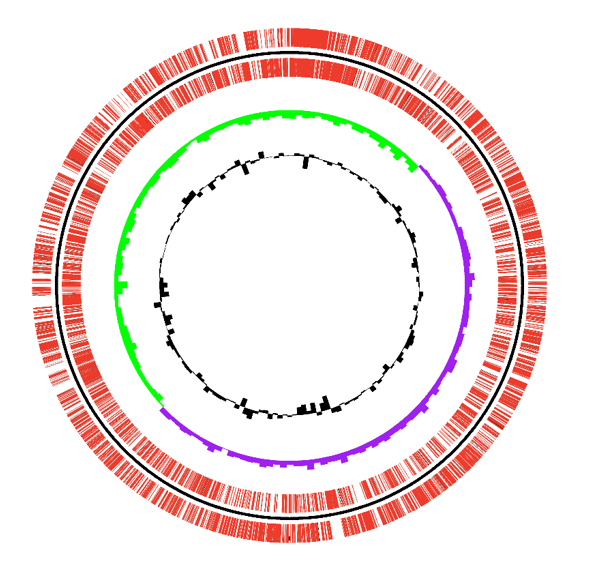
In order to visually display the results of gene annotation, we made a doughnut diagram of the genome by CIRCOS. It includes GC content, GC offset, positive and negative strand genes, rRNA, tRNA prediction and other information.
CRISPR sequence prediction
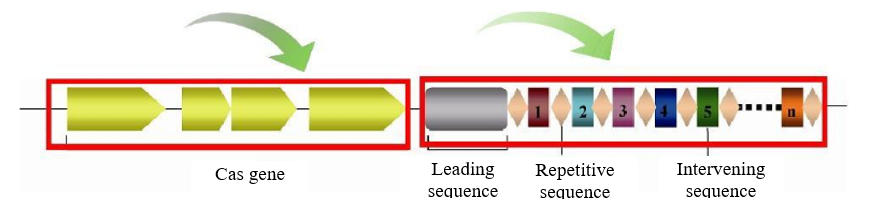
CRISPR is composed of repeat sequences and spacer sequences with similar length. It is the central component of the bacterial immune system and is responsible for sequence recognition. It also plays an important role in the bacterial immune system. It can resist the invasion of foreign DNA such as phage and plasmid. We use CRT to predict CRISPR sequence. The results could not only locate the position of CRISPR on the genome, but also include repeat sequences and spacer sequences.
COG notes
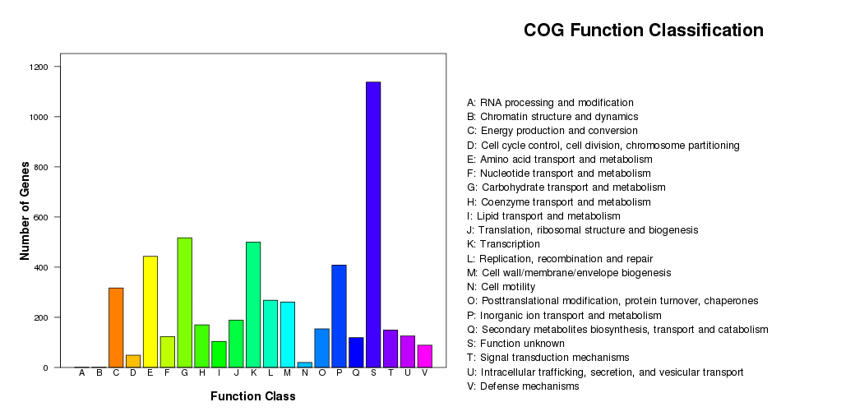
COG (Cluster of Orthologous Groups of proteins) is an NCBI classified database based on gene lineal homology. Combined with evolutionary relationships, COG classifies homologous genes from different species into different ortholog clusters. Currently, the COG has 4873 categories. Genes from the same ortholog have the same function, so functional annotations can be directly passed to other members of the same COG cluster.
-
Sort by
-
Date
Date(
)
Date
Date(
)
Impact Factor
IF(
)
Impact Factor
IF(
)



 Inquire
Inquire