Catalog Number|Packaging
Mat. No |
Ref. No |
No. of preps |
-
Description
Human Whole-genome sequencing (hWGS), mainly relying on high-throughput sequencing technique, obtains the whole genome sequence information of different human tissue samples, and comprehensively obtains the individual or group level structural variation information through bioinformatics analysis such as map reads to the reference genomes. hWGS provides important information for the study of pathogenesis and genetic mechanisms of diseases andcancers, and is also an important means on anthropogenesis and population evolution.
-
Technical Workflow
Experiment design → Sample preparation → Library construction & sequencing → Data analysis → After-sales service
-
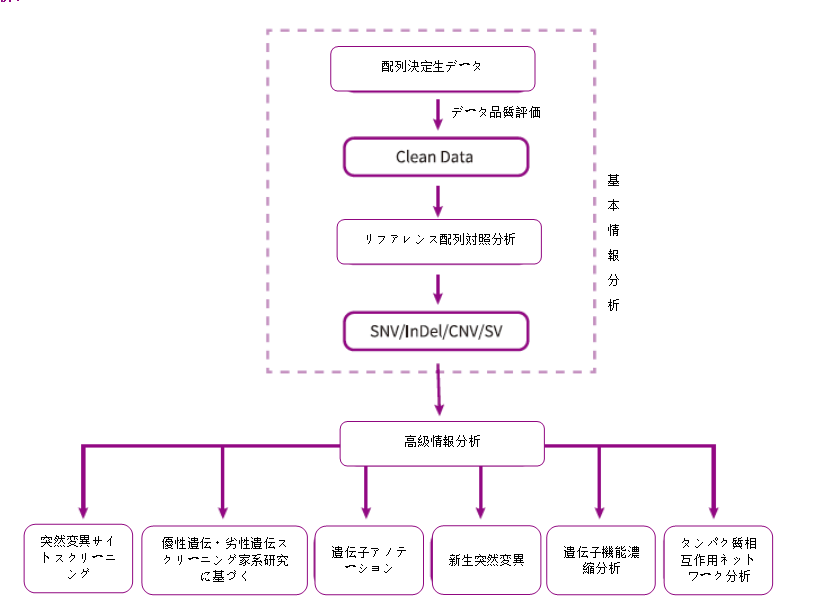
Analysis Workflow
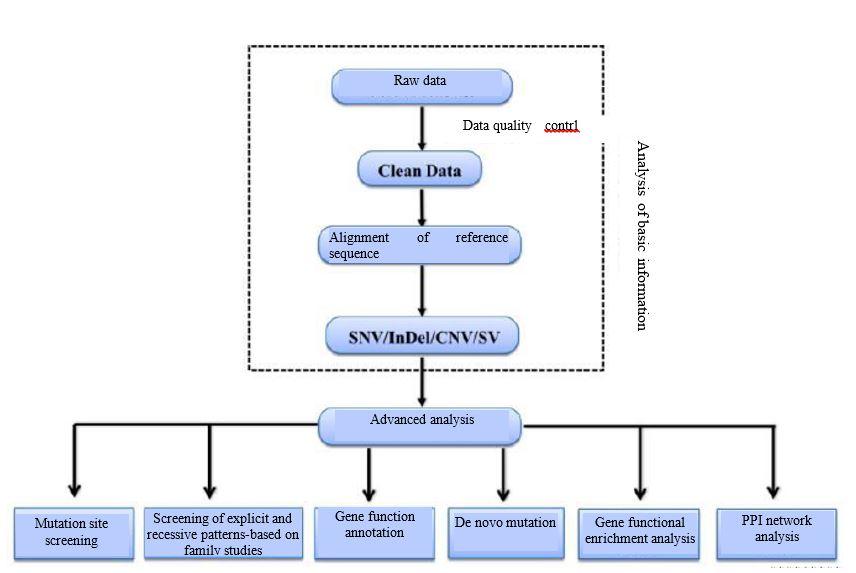
-
Features
1. Obtain comprehensive genetics variations;
2. Provide various advanced analysis according to different needs;
3. Provide multiple database support, such as TCGA, IPA, HGMD, IVA and so on;
-
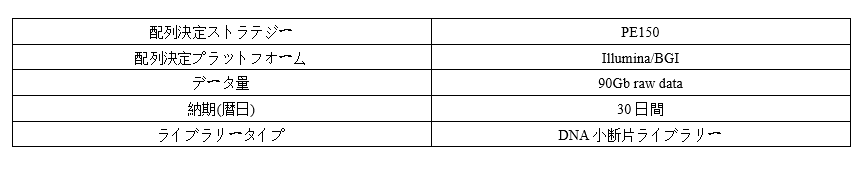
Parameter

-
Applications
Pathogenesis, Genetic mechanism, biomarkers, Cancer research
-
Requirements of Samples
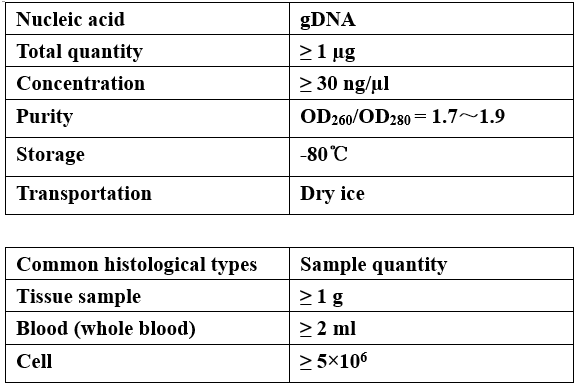
-
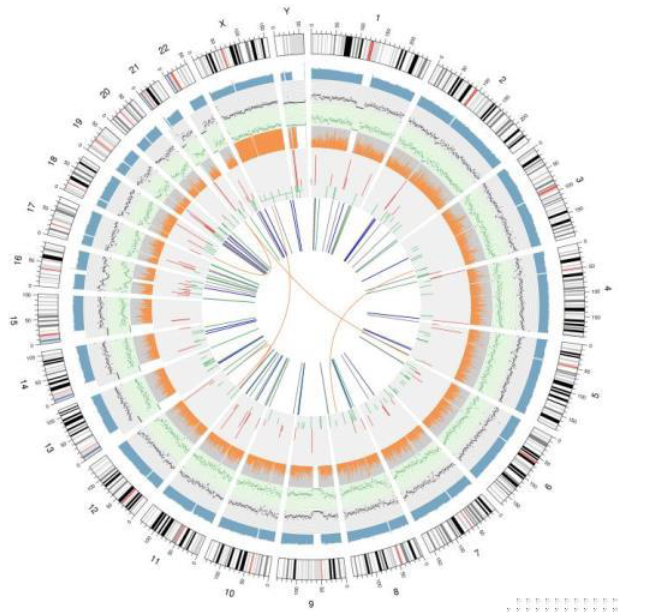
Results Demo
Circos based on whole genomic results
Candidate gene GO: Biological Process enrichment bar plot
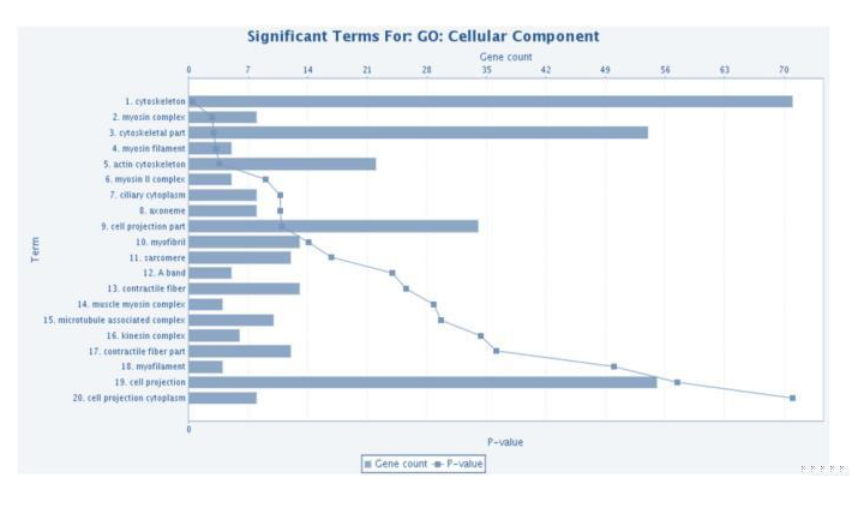
The candidate genes obtained were carried out functional enrichment such as GO, Pathway, miRNA, Disease (Chen J et al. 2009-2), Proteome, Phenotype, etc. by online tool ToppGene Suit-ToppFun. In enrichment analysis, the multiple test correction value Q-value of P-value is used to control FDR.
In addition, more stringent multiple correction method Bonferroni can also be adopted. The default enrichment significance threshold is Q-value < 0.05 that controls FDR.
PPI network analysis of candidate gene set
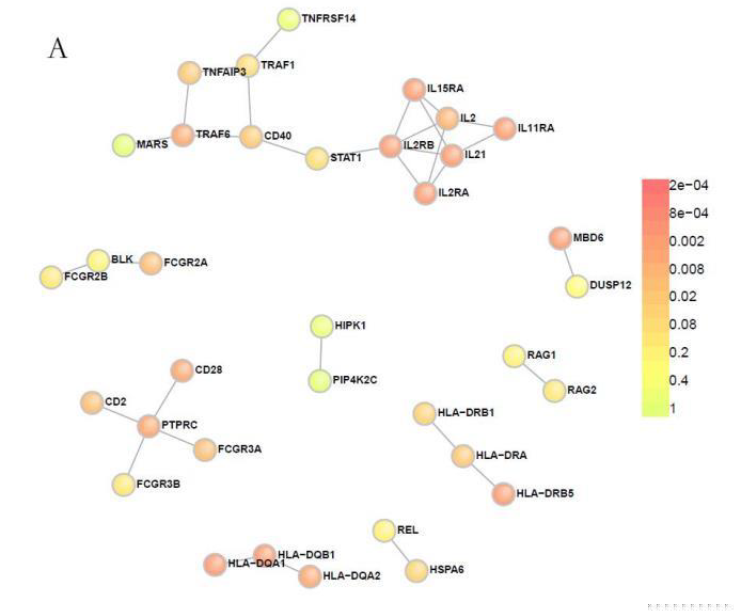
PPI network analysis was carried out on the candidate gene set to find the co-regulated genes that interact. Similar to metabolic pathways, mutant genes may differ from patient to patient, but phenotypes caused by interacting gene mutations may be consistent. Using the online tool DAPPLE to look for mutant genes that interact with each other can help locate virulence genes that are truly related to diseases.
-
Sort by
-
Date
Date(
)
Date
Date(
)
Impact Factor
IF(
)
Impact Factor
IF(
)



 Inquire
Inquire