Catalog Number|Packaging
Mat. No |
Ref. No |
No. of preps |
|
4995267 |
GNR112-01 | 24 rxn |
|
4995268 |
GNR112-02 | 96 rxn |
-
Storage
The kit should be stored at -15~-30°C. -
Description
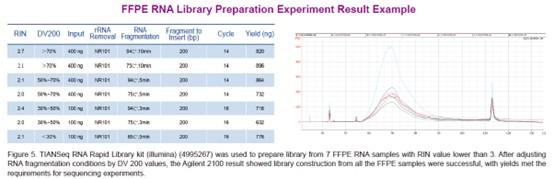
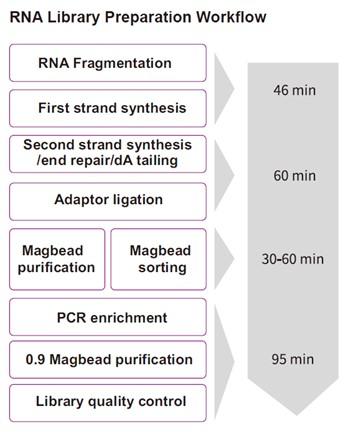
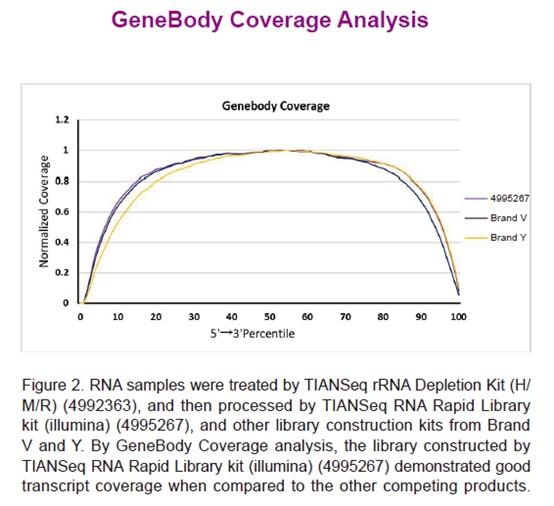
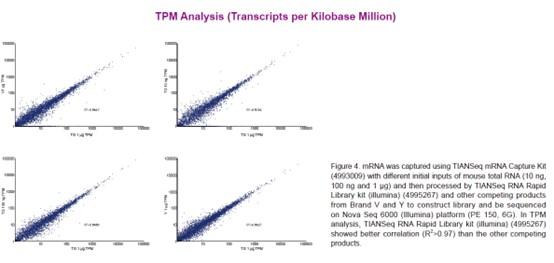
TIANSeq RNA Rapid Library kit (illumina) is a non-stranded transcriptome library construction kit specifically developed for Illumina high-throughput sequencing platform. The kit adopts a rapid one-tube process for rapid library construction from RNA samples. After the first strand synthesis, double strand cDNA synthesis, end repair and dA tailing can be completed in one step. And the reaction products can be used for adaptor ligation directly, without purification. In addition, the kit uses a specially designed high-fidelity polymerase, which provides PCR amplified products of high fidelity without base preferences.
The starting sample should be total RNA with rRNA excluded, (mRNA and other non-coding RNA are retained), or mRNA obtained by direct isolation from total RNA. Sample input ranges from 10 ng to 1ug for total RNA samples, input of mRNA sample could be as low as 500 pg.
-
Features
■Suitable for transcriptomic analysis of mRNA, and other non-coding RNA (e.g., lncRNA), except from rRNA.■ Easy protocol for rapid library construction.
■ High library conversion rate for effi cient conversion from verylow sample input amount.
■ The PCR amplifi cation is of high fi delity and free of base preferences.
-
Applications
It is applicable to the construction of RNA library of Illumina high-throughput sequencing platform. Applicable sample input: 10 ng ~1 μg of total RNA; As low as 500 pg of initial mRNA of animals, plants, fungi, etc.


-
Sort by
-
Date
Date(
)
Date
Date(
)
Impact Factor
IF(
)
Impact Factor
IF(
)



 Inquire
Inquire